Protocol for tissue clearing and lightsheet imaging
1. Tissue clearing
PEGASOS tissue clearing technique, developed by the Zhao lab, enables the simultaneous clearing of both soft and hard tissues, making it suitable for 3D fluorescence imaging of whole organs. In our study, we applied the PEGASOS protocol to clear whole-brain samples from double-transgenic mice (Tek-Cre; Ai47) in which endothelial cells were labeled with a fluorescent reporter. This was done in preparation for subsequent 3D whole-organ imaging using light-sheet microscopy. The tissue clearing procedure for mouse brains was performed as follows:
- Mice were anesthetized and transcardially perfused as previously described for immunohistochemical experiments.
- Brain tissues were fixed in 4% paraformaldehyde (PFA) for 2 hours, followed by three 30-minute washes in PBS at 4°C with shaking at 80 rpm.
- Samples were then decolorized by incubation in 25% ethylenediamine (Quadrol) for 1 day.
- For delipidation and gradient dehydration, samples were sequentially submerged in 30%, 50%, and 70% tertiary butanol for 6 hours each at 37°C with constant shaking.
- Dehydration was completed by incubating the samples for 2 days in tB-Quadrol solution (80% tertiary butanol and 20% Quadrol), with the solution replaced daily.
- The final clearing step involved incubation in BB-BED468 solution (BB:BED468 at a 3:2 ratio) for at least 24 hours, or until the tissue became transparent.
- The clarified samples were stored and imaged in BB-PEG clearing medium at room temperature (Fig. 1).

Figure 1 | PEGASOS-based tissue clearing of mouse brain samples. (a) Schematic of the PEGASOS soft tissue clearing protocol. (b) Representative images of a mouse brain sample at various stages of the clearing protocol. (1) The sample after the dehydration step. (2) The sample submerged in the clearing solution. (3) The cleared sample after 24 hours, with ruler for scale. (4) The sample submerged in the refractive index-matched imaging medium before light-sheet microscopy. Scale bar, 5 mm.
2. Light sheet fluorescence imaging
Tiled light-sheet microscopy was performed using a Nuohai LS18 microscope (Nuohai Biological Scientific Instruments Co., Ltd., Shanghai). The entire brain was imaged in dual-side illumination mode with an excitation wavelength of 488 nm (Fig. 2). Imaging parameters, including tiling overlaps (X=31%, Y=19%, L&R=31%), are detailed in Table 1. A 6.3× objective was used for all acquisitions.
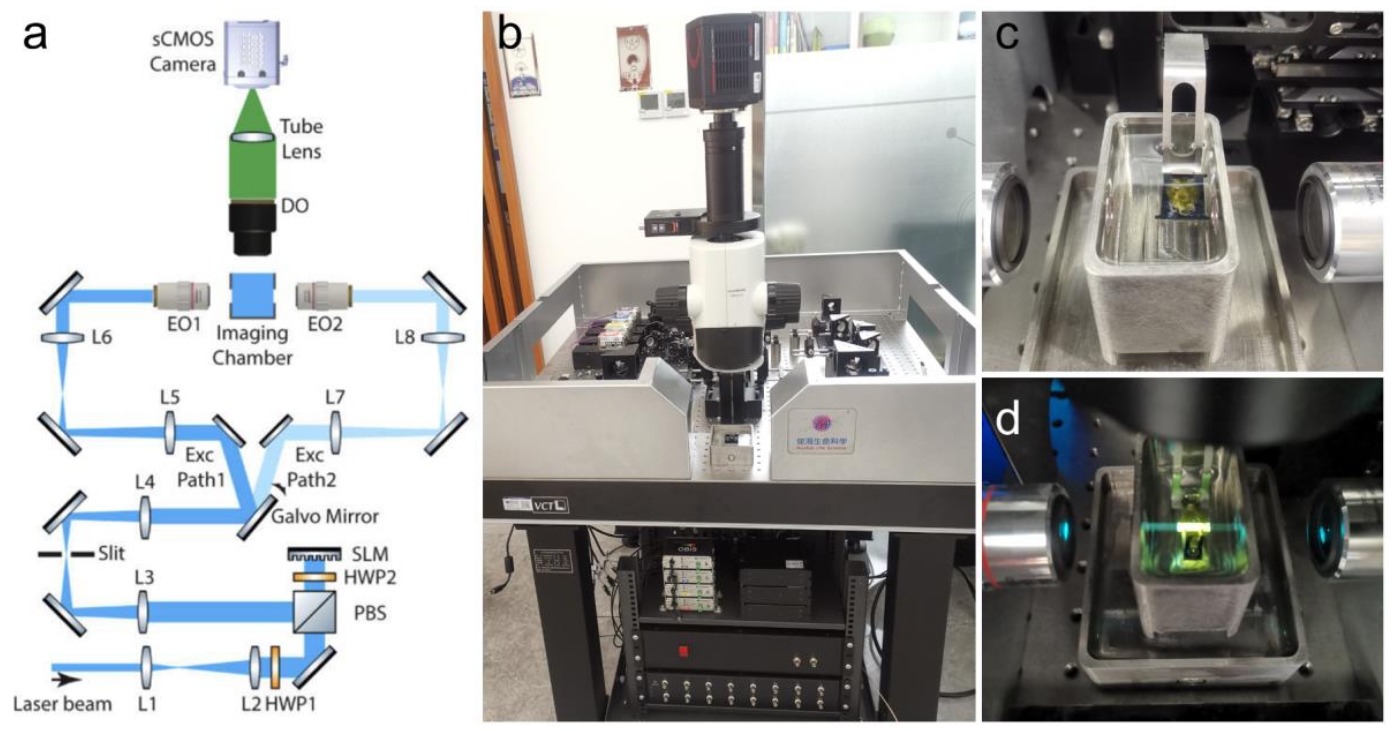
Figure 2 | Light-sheet microscopy imaging of cleared mouse brain samples. (a) Schematic of the optical path of a tiled light-sheet microscope. (b) Photograph of the tiled light-sheet microscope (Nuohai LS18). (c) The process of mounting a cleared mouse brain sample. (d) Diagram of the imaging process: excitation light is emitted from the illumination objectives on both sides, while a single objective at the top focuses and collects the emitted fluorescence.
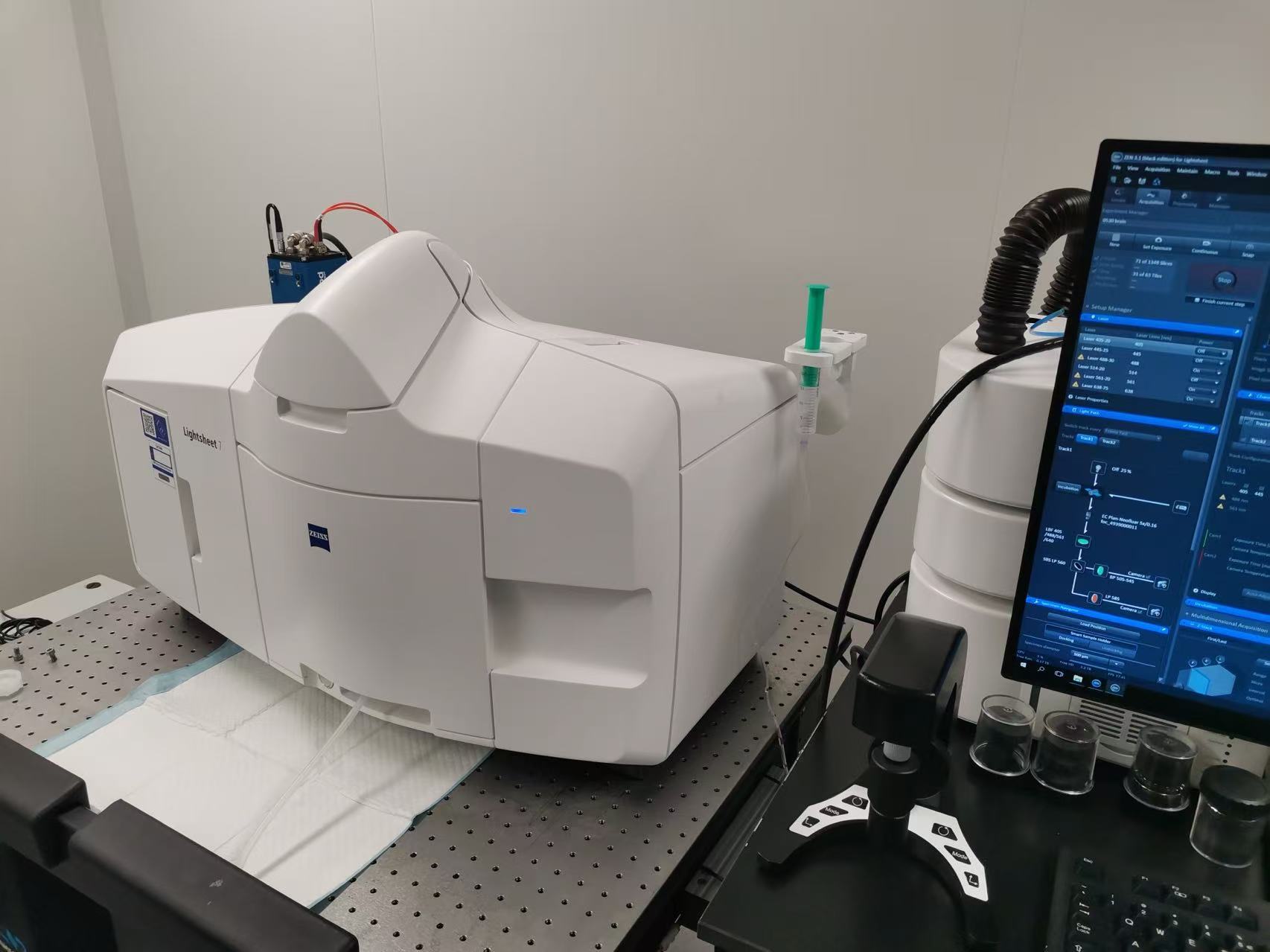
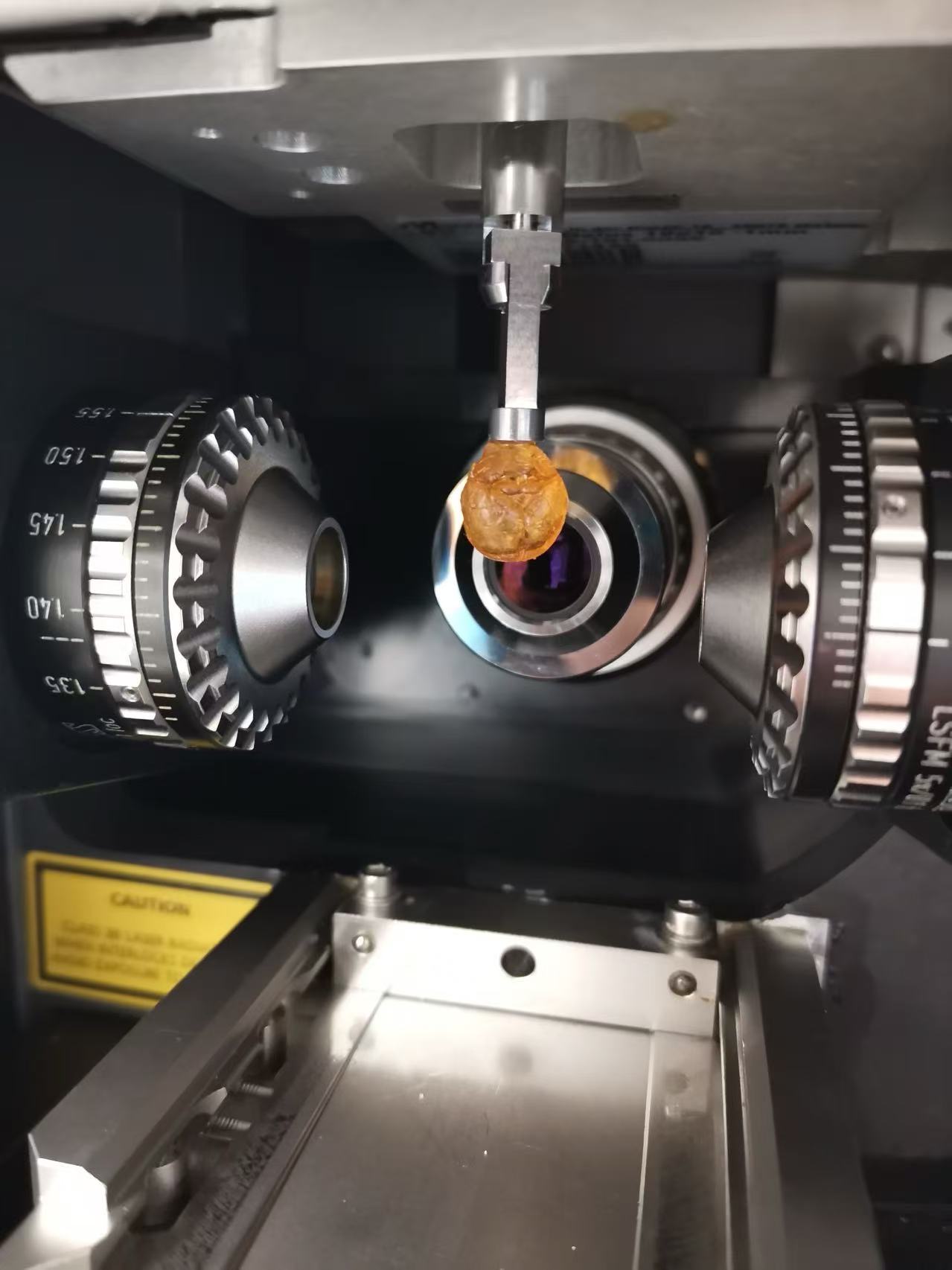


Figure 3 | Imaging of cleared mouse brain sample with ZEISS Lightsheet 7.
3. Data analysis and results
Cleared brain samples from normal adult Tek-Cre;Ai47 mice were acquired on the Nuohai LS18 (Fig. 2). For the Braf-Ai14-AVM mouse model, cleared brain samples were imaged on a conventional light-sheet microscope (Lavision UltrascopeII) with the assistance of the imaging core facility at the Chinese Institute for Brain Research (CIBR). Imaging results from these mice are shown in Fig. 4.
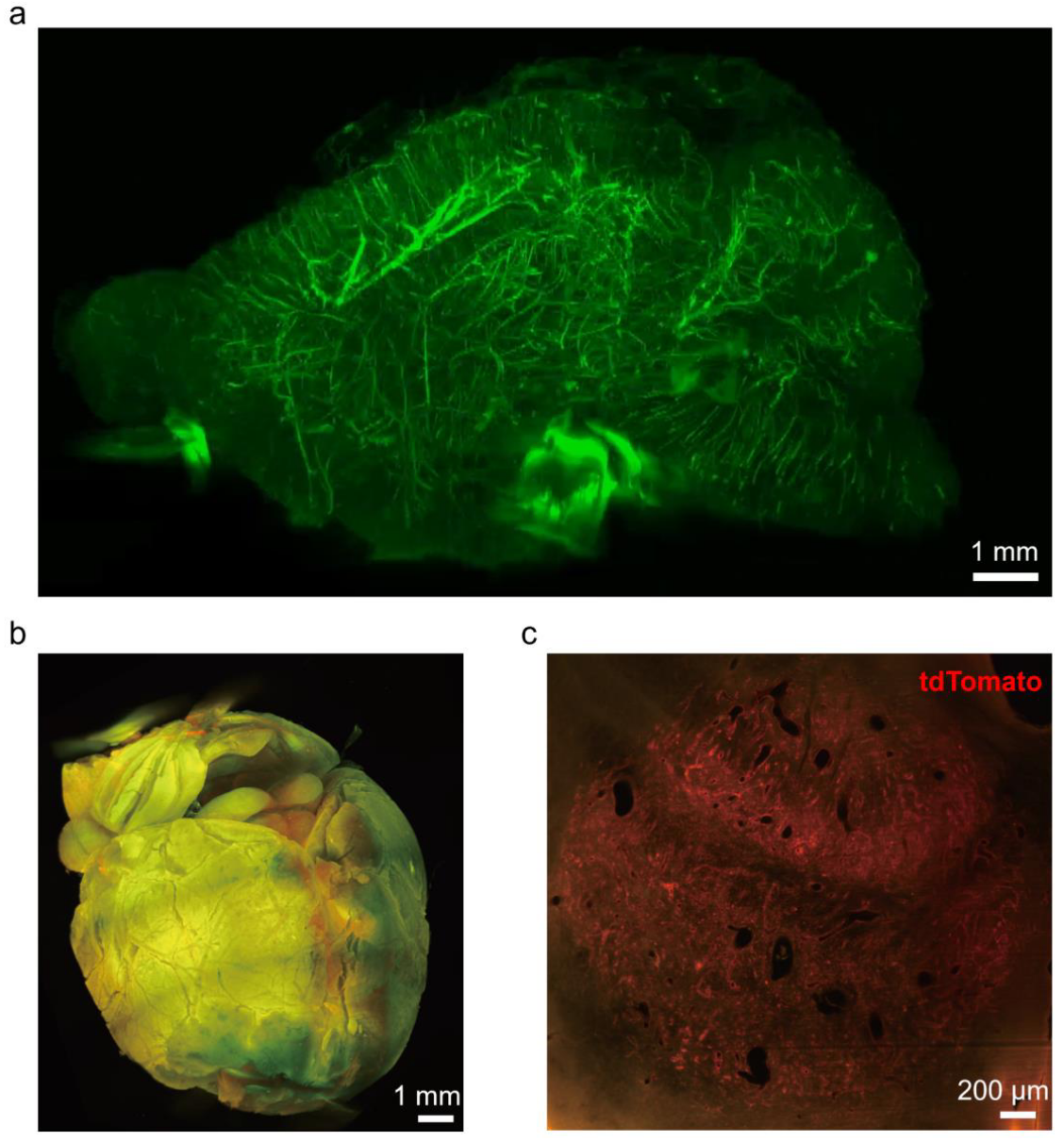
Figure 4 | 3D whole-brain imaging of normal and Braf-AVM disease model mice. (a) 3D reconstruction of a cleared brain from a normal adult Tek-Cre; Ai47 transgenic mouse. (b) 3D reconstruction of a cleared brain from a Braf-AVM model mouse. The sample was generated by in situ injection of AAV-PHP.eB-mPro723-Cre virus into a Braf-CA; Ai14 transgenic mouse to induce AVM, followed by tissue clearing. (c) Magnified cross-section through the center of the spherical lesion shown in (b). The tdTomato fluorescence signal indicates endothelial cells with the induced Braf point mutation.
Softwares for lightsheet imaging data analysis:
- BigStitcher [PMID: 31384047 | DOI]
The BigStitcher is a software package that allows simple and efficient alignment of multi-tile and multi-angle image datasets, for example acquired by lightsheet, widefield or confocal microscopes. The software supports images of almost arbitrary size ranging from very small images up to volumes in the range of many terabytes, which are for example produced when acquiring cleared tissue samples with lightsheet microscopy.
Tutorials:
- Reconstructing terabyte sized light sheet datasets
