Nutrient sensing mTORC1 pathway
Nutrient sensing is a fundamental biological process that enables organisms to adapt to their environment by regulating cell growth and metabolism in response to nutrient availability. mTOR signaling pathway is evolved to be able to sense environmental cues and coordinates cellular growth, we are focusing the question: How does mTOR sense nutritional cues and coordinates cellular growth ?
One the road of discovering new nutrient sensors……
Introduction
Nutrient sensing is a fundamental biological process that enables organisms to adapt to their environment by regulating cell growth and metabolism in response to nutrient availability [1]. A key pathway in this process is the mechanistic target of rapamycin complex 1 (mTORC1), which coordinates cellular responses to amino acid and glucose levels. Despite significant advances in understanding mTORC1 signaling, only a limited number of nutrient sensors has been discovered. The identification of sensors like Sestrin2 [2], CASTOR1 [3], LYCHOS [4], SAMTOR [5] and Unmet [6] has provided insights into how cells detect nutrients such as leucine, arginine, cholesterol, and S-adenosylmethionine. However, the evolutionary diversity of nutrient sensing mechanisms suggests that there are likely many undiscovered sensors [7], particularly in non-mammalian organisms occupying unique ecological niches. Identification of such sensors should reveal novel regulatory components and mechanisms of the mTORC1 pathway.
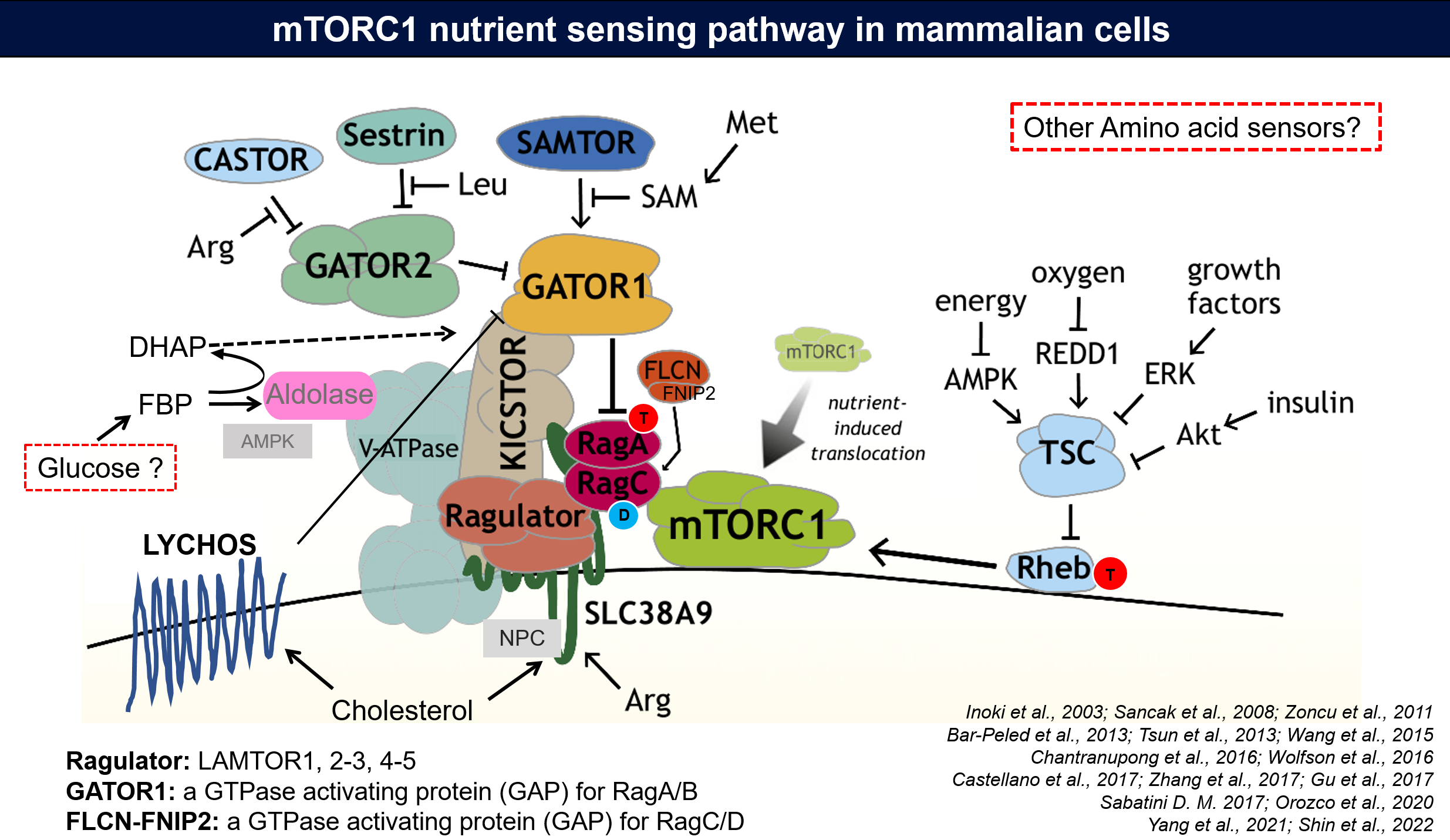
To be active, mTORC1 requires the environmental presence of amino acids and glucose. While a mechanistic understanding of amino acid sensing by mTORC1 is emerging, how glucose activates mTORC1 remains mysterious. It was previously reported that mTORC1 senses a metabolite downstream of the aldolase and upstream of the GAPDH-catalysed steps of glycolysis and pinpoint dihydroxyacetone phosphate (DHAP) as the key molecule [8]. Now, we are also on the way to discover the glucose sensor.
Our discoveries may lead to new treatments of cancer, aging and many other diseases.
References
- Chantranupong, L., Wolfson, R. L. & Sabatini, D. M. Nutrient-sensing mechanisms across evolution. Cell 161, 67-83 (2015).
- Wolfson, R. L. et al. Sestrin2 is a leucine sensor for the mTORC1 pathway. Science 351, 43-48 (2016).
- Chantranupong, L. et al. The CASTOR Proteins Are Arginine Sensors for the mTORC1 Pathway. Cell 165, 153-164 (2016).
- Shin, H. R. et al. Lysosomal GPCR-like protein LYCHOS signals cholesterol sufficiency to mTORC1. Science 377, 1290-1298 (2022).
- Gu, X. et al. SAMTOR is an S-adenosylmethionine sensor for the mTORC1 pathway. Science 358, 813-818 (2017).
- Liu, G. Y., Jouandin, P., Bahng, R. E., Perrimon, N. & Sabatini, D. M. An evolutionary mechanism to assimilate new nutrient sensors into the mTORC1 pathway. Nature Communications 15, 2517 (2024).
- Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302-310 (2015).
- Orozco, J.M., Krawczyk, P.A., Scaria, S.M. et al. Dihydroxyacetone phosphate signals glucose availability to mTORC1. Nat Metab 2, 893–901 (2020).
